- About
- Organization
- Organization Overview
- Dean’s Office
- Department of Bioengineering and Therapeutic Sciences
- Department of Clinical Pharmacy
- Department of Pharmaceutical Chemistry
- Quantitative Biosciences Institute
- Org Chart
- Research
- Education
- Patient Care
- People
- News
- Events
Computer models to predict drug clearance by liver cells show promise
By David Jacobson / Tue Nov 22, 2011
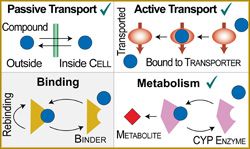
A schematic diagram of the mechanisms built into each cell in the in silico hepatocyte cultures. Checkmarks indicate events affected by the physical and chemical properties of a specific virtual drug object. For example, depending on such properties a given drug bound to an enzyme can have a different probability of being metabolized or released. (Source: Drug Metab Dispos. 2011 Oct;39(10):p. 1910-20)
A holy grail of drug discovery is to answer key questions about potential new drugs less by experiments in petri dishes and lab animals and more by faster, cheaper engineering efforts using predictive computer models.
UCSF School of Pharmacy faculty member and bioengineer Anthony Hunt, PhD, and colleagues are creating computer programs that simulate the in vitro (i.e. laboratory) interactions between drugs and cultures of hepatocytes—cells that make up most of the liver and metabolize drugs.
A key goal of these in silico (i.e. computer simulated) hepatocyte cultures (ISHCs) is to be able to predict the rate, relative to known drugs, at which a potential new drug will be cleared (i.e. removed) from the blood by the liver. Drugs that are cleared too fast may be ineffective against disease, while drugs cleared too slowly might accumulate to toxic levels.
Discovering this information more quickly means researchers will not waste time and resources developing drugs that would fail to work in vivo (i.e. in the body) or may be able to address clearance issues sooner by altering a drug’s chemistry.
A study published in October 2011 and co-authored by Hunt explored the feasibility of using ISHCs to predict clearance by comparing the results of their simulated interaction with dozens of drugs for which in vitro clearance rates are known.

C. Anthony Hunt, PhD
About this Article
UCSF researchers: Anthony Hunt, PhD, is a faculty member of the Department of Bioengineering and Therapeutic Sciences, a joint department of the UCSF Schools of Pharmacy and Medicine. The study was co-authored by Shahab Sheikh-Bahaei, PhD, a recent graduate of the UCSF/UCB Joint Graduate Group in Bioengineering.
Journal citation: Sheikh-Bahaei S and Hunt CA, “Enabling clearance predictions to emerge from in silico actions of quasi-autonomous hepatocyte components.” Drug Metabolism and Disposition, Vol. 39, Issue 10. P. 1910-1920 (October, 2011). Epub July 18, 2011.
The research
Hunt and Sheikh-Bahaei created computer programs that simulate hepatocyte cultures. Their greatly simplified virtual liver cells included key drug metabolizing features such as transporters (membrane-based proteins that control the entry and exit of drugs) and enzymes (modeled in part after cytochrome P450 enzymes that are commonly involved in drug metabolism).
The scientists also:
- Created “drug objects” that acted as simulation stand-ins for the molecules of real-world drugs ranging from caffeine to morphine and from the antacid omeprazole to the blood-thinner warfarin. These bore “name tags” that included clearance-relevant physical and chemical properties.
- Assigned “time steps” to the binding and release of drugs by transporters and enzymes, and probabilities to their metabolism by the latter.
- Established rules for how certain physical and chemical properties of a drug molecule (i.e. size, charges, structures) would influence the probability of a given event in an ISHC.
Next came hundreds of simulation experiments—putting 8,000 virtual drug molecules into in silico cultures made up of 1,000 hepatocyte cells.
Based on how well the simulation clearance results matched those of actual in vitro experiments, the mechanisms of the in silico hepatocytes (e.g. the ratio of transporters to enzymes) and the impact of certain characteristics of drug molecules on intracellular transport, binding, and metabolism were modified to increase the model’s accuracy.
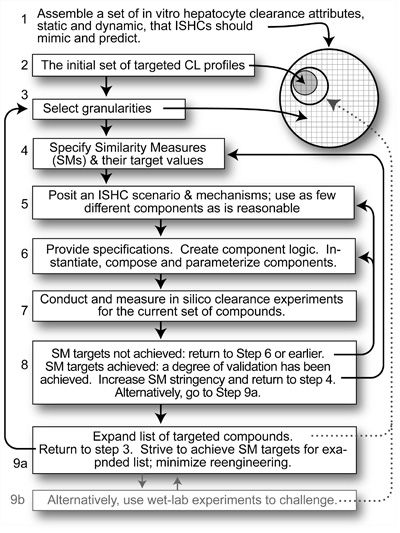
The process used to systematically develop and improve an in silico hepatocyte culture.
The results
The scientists ran simulations to determine the clearance rates when their ISHCs were exposed to virtual doses of 73 different drugs—all those for which similarly measured in vitro human hepatocyte clearance rates have been reported.
For 39 of the drugs, a full range of hepatocyte culture clearance rates have been established. Among those drugs, 79 percent of the clearance rates predicted by the ISHCs were within those real-world ranges of distribution. That is, they were within two standard deviations, accounting for 95 percent of all reported clearance rates. This exceeded the researchers’ goal of having at least half their model’s rate predictions fall within that range.
By comparison, a method that attempts to predict hepatocyte drug clearance by correlating a drug’s physical and chemical properties with the data for clearance of similar drugs (i.e. multiple linear regression) was accurate within two standard deviations only 23 percent of the time.
As the scientists noted, “Thus, feasibility was demonstrated: the concrete detail built into ISHCs enabled acceptable performance.”
The predictive accuracy of both computer model and statistical correlation methods for the full set of 73 drugs “were similar; both were poor”—with 38 percent and 32 percent respectively within a factor of two (i.e. twice as much or half as much) of the available clearance values.
In fact, that lesser accuracy may simply reflect that for 34 of the drugs only a single in vitro clearance rate was available. And those rates can vary widely due to laboratory methods or differences among the clearance rates of cells that come from different people.
Still the scientists concede, “Considerable refinement is needed to make ISHC predictions useful for supporting research and development decision making.”
But they also assert that repeated refinement of the mechanisms being modeled “provides a means of achieving improvement by making ISHCs incrementally more complicated or fine-grained.”
In other words, adding more properties of drug molecules and more mechanistic details of their metabolism by liver cells will make the currently simple computer models—which are already better than mere statistical correlation—even more accurate.
The implications
This research by the Hunt Lab is another step toward physiologically-based pharmacokinetic modeling, which seeks to predict the absorption, distribution, metabolism, and excretion of drugs by creating mathematical models of how they are thought to be processed by the body.
As Hunt and Sheikh-Bahaei write: “We envision plugging hepatocytes into validated in silico liver analogs within larger patient analogs.”
The scientists think simulations will be able to predict relative drug clearances more accurately as further details of the interaction between liver metabolism and the physical and chemical properties of drugs are discovered and built into their computer models.
In addition, computer models of liver cell cultures will allow scientists to investigate why different drugs are cleared at different rates and the reasons for variations in clearance among different patients.
Eventually such computer models could also be used to better anticipate influences of genetic differences, drug interactions or liver diseases on a patient’s drug clearance, generating personalized dosing for a particular patient.
Tags
Keywords:
Category:
Sites:
School of Pharmacy, Department of Bioengineering and Therapeutic Sciences, PharmD Degree Program, QBC, PSPG
About the School: The UCSF School of Pharmacy aims to solve the most pressing health care problems and strives to ensure that each patient receives the safest, most effective treatments. Our discoveries seed the development of novel therapies, and our researchers consistently lead the nation in NIH funding. The School’s doctor of pharmacy (PharmD) degree program, with its unique emphasis on scientific thinking, prepares students to be critical thinkers and leaders in their field.




